HUPO ECR Online Panel Discussion - Getting recognised for your work
The HUPO Early Career Researcher (ECR) committee has organised a discussion panel on Getting recognised for your work and have asked me to participate - thank you! I am always keen on such events, organised by and for ECRs.
Introductions
The first part of the panel is a short 5-minute introduction of the panellists, including Prof Stacy Malaker from Yale University and Dr Juan Antonio Vizcaino from the EMBL-EBI and myself.
I prepared this career flow chart as a visual aid:
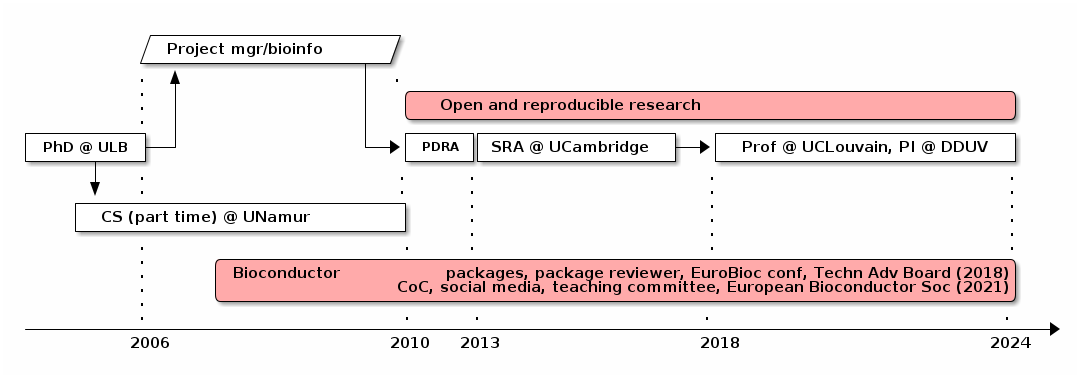
- I earned my PhD in 2006, from the Free University of Brussels (ULB). My PhD work focused on the evaluation of different types of evolutionary genetic markers to study cetaceans phylogeny.
- During my PhD (probably around 2004 or so), when my work and interests shifted toward bioinformatics, I started a part-time degree in computer science at the University of Namur. That lasted until I left Belgium for the UK in 2010, after completing all my exams, but before finishing my masters project - I never graduated.
- After my PhD, I worked for 3 years in industry, in a small company (we probably were about 15 employees). I didn’t see much point in continuing in academia at that point, considering my experience so far and my personal situation. The environment and general atmosphere was very much like an academic lab, with many more collaborations within the team, and clear and common objectives. This goal-oriented work environment was a very refreshing experience that has been influential for the next steps of my career. At some point, I felt I was starting to run in circles and got a chance to move back to academia, at the University of Cambridge nonetheless.
- In 2010, I started a post-doctoral research associate (PDRA) position in the Cambridge Centre for Proteomics, working on mass spectrometry-based proteomics.
- In 2013, I got promoted to senior research associate (SRA), which allowed me to earn some grants as main PI and develop a small research team.
- In 2018, I joined the UCLouvain as a professor of bioinformatics. I teach in the faculty of pharmacy and biomedical sciences (FASB) and run the CBIO computational research group in the de Duve Institute.
An interesting fact is that I started working on DNA during my PhD, moved on with RNA in the private company, and since moving back to academia, I have been focusing on proteins: my career followed the main path of the central dogma of molecular biology.
To give more context to my career path, I also highlight some other activities and interests, that have guided and supported my academic activities.
-
I started to realise the importance of open and reproducible research around 2010, both with respect to the rigour of doing research, but also in the light of the (at times) oppressive and restrictive global research environment ECR have to endure. The desire for others to benefit from my research by making it as open, collaborative and reproducible as possible, and being vocal about it, has followed me since then.
-
The Bioconductor project has been instrumental for me. It has allowed me over the years to meet and be influenced by outstanding scientists, and has offered an international environment in which I was able to grow and flourish. I published my first (now retired) Bioconductor package around 2007 (Bioconductor 2.2 and R 2.7), and many more followed. I am a Bioconductor package reviewer, a member of the European Bioconductor (EuroBioc) conference organisation committee (I was a local organiser for a handful EuroBioc conferences in Cambridge, in 2019 in Brussels, and in 2023 in Ghent), have been part until recently of the Code of Conduct committee, am part of the social media working group, I co-lead the Teaching committee, am since 2018 member of the Technical Advisory Board, and co-created, in 2021, the European Bioconductor Society.
Recognition
The panellists were also asked to comment on how they have been recognised for their work, with a particular emphasis their time as ECRs. This is of course very subjective, and I’m not sure if my answer will reflect how I have been recognised (assuming I have), or how I hope I have been. I will also try to look beyond papers, the obvious academic outputs - without those, there is little chance to get academic recognition. This is of course a major problem, as there’s much more to research than papers:
An article about computational science in a scientific publication is not the scholarship itself, it is merely advertising of the scholarship. The actual scholarship is the complete software development environment and that complete set of instructions that generated the figures.
[Buckheit and Donoho 1995, after Claerbout]
I think I’m known for the computational development and applications in spatial (2010…) and single-cell proteomics (2018…), my efforts to produce open and reproducible research, open and collaborative software development, my R/Bioconductor contributions (some packages have been around for since 2010) as well as my involvement in teaching, such as international workshops (for example the mythical - for me at least - Bioconductor CSAMA course workshop).
One noteworthy aspect of my publication strategy, that highlights my efforts for openness and reproducibility, is the workflow that typically starts with the release of the software (more often than not after review, on Bioconductor), then the publication of a pre-print with code to reproduce the analyses and, eventually, a peer-reviewed paper.

I think I have gained some reputation as someome having expertise in computational quantitative proteomics, including demonstrable technical skills, in addition to more standard scientific/academic output.
In terms of recognition, I suppose that invitations to give talks (for scientific outputs), teach at workshops (pedagogical and technical skills) and to submit papers are obvious goals. Being recognised for my open and collaborative contributions with a Bioconductor community award is one of my proudest moments. High on the list are also the many contributions that the MSnbase package benefited from - some of these indirectly initiated the collaborations that lead to the creation of the R for Mass Spectrometry initiative.
But what matters the most, in my eyes, and what in the end is the most meaningul recognition, are the (shared) values that we promote with the research we do, and the intrinsic motivation that drive us.
Questions
We were also asked to prepare answers to three short questions. These have been pre-determined by the HUPO ECR to get things going and give us, the panellists, a chance to think about the comments we would like to make.
When hiring a new postdoctoral researcher for your group, what are the most important attributes for them to have on their CV? What do you look for other than publication history?
Here are a couple of things I look for, and that I consider absolutely essential, much more important than papers. Papers are only one of the attributes that will help me assess the following:
- Does the candidate’s skills match the project’s needs?
- What are concrete signs of mastery? I perform regular (and constructive) appraisals with the researchers in my group, and one question in that appraisal is “What do you want to become an expert in?”. In a CV, I want to find what the candidate is an expert in, whey they can teach me/bring to the lab.
- I also need to see public/open
code, such as for example active Github/Gitlab profiles and repositories and contributions (to their or other’s code base).
And of course, last but not least, will the person be a good lab/team member? It is of course very difficult (and arguably subjective) to assess, but we (the lab) will be attentive to red flags pointing to the contrary. In case of doubt, I will invite the candidate on site if the interview (always with the whole group) was remote. I wouldn’t want to take any risks that could harm the cohesion and well-being of the group.
How would you recommend that ECRs promote their work other than research e.g., teaching, outreach, committee work? Is there anything they can do other than add a line on their CV?
- Yes, ‘add lines’ to your CV, but not at all costs. Be pragmatic! Not need to run or teach workshops several time a year to demonstrate that you have done some teaching. Don’t forget that your post-doc years should be the most productive research-wise of your career!
- Promote your research, don’t be a vehicle for some else’s research (typically your advisor), don’t limit yourself to merely doing it. Show how you go the extra mile - for example by delivering reproducible research.
- Do things you like! Nothing beats motivation when it comes to convincing others that you are good at what you do.
How much does networking (either via social media or in-person meetings) play a role in promoting your work?
It is very important! Networking are opportunities to learn, share, discuss, and make yourself known, … Networking can be hard though, so don’t be too hard on yourselves. It takes time.
Here’s a simple example illustrating the importance of building a network: I happily spare myself organising and running interviews when I can find a candidate in my direct or indirect network.
To build that network, there’s of course the in-person or remote conferences and workshops, there may be social media (might not be for everybody), but also Github issues and code and documentation contributions and typo fixes. The large and small contributions are very concrete examples that address the first question above.

